武汉华美生物工程有限公司CUSABIO®品牌商
15 年
手机商铺
- NaN
- 0
- 0
- 2
- 2
推荐产品
公司新闻/正文
B2M/β2微球蛋白:从抗原呈递核心到免疫治疗与疾病预测的关键靶点
456 人阅读发布时间:2026-03-16 13:29
β2微球蛋白(B2M)作为MHC-I分子的轻链亚基,是内源性抗原呈递通路的核心组分,在CD8+ T细胞识别病毒与肿瘤抗原过程中发挥不可替代的作用。除经典免疫功能外,B2M还参与神经认知调控、代谢稳态维持等非免疫生理过程,其表达异常与肿瘤免疫逃逸、自身免疫病、神经退行性疾病及心血管风险密切相关。近年来,B2M作为免疫治疗响应预测标志物、通用型细胞治疗工程化靶点以及自身免疫病干预新靶点的转化价值日益凸显。本文系统梳理B2M的结构功能、调控机制、疾病关联及靶向治疗进展,为相关研究与临床转化提供参考框架。
1. B2M的结构特征与MHC-I轻链功能:抗原呈递的分子基础
2. 可溶性B2M的非免疫效应与生物标志物价值:优势与局限
3. B2M-MHC-I抗原呈递机制与调控:从装配到免疫逃逸
4. B2M相关信号通路:免疫细胞互作与肿瘤内源性轴
5. B2M与疾病:免疫逃逸、微环境、监测与非肿瘤病理
6. B2M靶向药物最近研究进展
7. B2M研究工具推荐:重组蛋白、抗体与ELISA试剂盒选型指南
1. B2M的结构特征与MHC-I轻链功能:抗原呈递的分子基础
B2M最早被定义为MHC-I分子轻链,其经典功能是与MHC-I重链(HC)非共价结合形成稳定复合体,维持肽槽结构并促进内源性肽的装配与呈递,从而支持CD8+ T细胞对病毒与肿瘤抗原的识别。相关工作亦从肽呈递预测的角度强调MHC结构域关键口袋位点与肽两端的重要性,侧面支持“结构稳定性—呈递命运”之间的内在关联 [1]。在进化与突变层面,B2M属于IgSF基本构件,其序列保守性与功能优化可由拓扑与动力学模型解释;同时也识别出如D76N等聚集易感突变,与淀粉样沉积现象相符,提示B2M兼具结构支架与潜在病理聚集倾向 [2]。
值得注意的是,近年证据提示MHC-I重链可在未结合B2M和/或肽的状态下以B2M-free HC形式出现在细胞表面,并进一步形成同/异二聚体;此类分子在炎症刺激或细胞激活后上调,糖基化模式与经典复合体不同,且可能与KIR受体、促生长因子及细胞因子等发生相互作用;在动物模型中,缺失B2M背景下HLA-B27诱发自发性关节病提示其可能影响免疫稳态与自身免疫表型 [3]。
总体而言,现阶段对B2M-free HC的发生学、在体功能及其与经典MHC-I通路效率之间的因果关系仍缺少系统的生化与结构证据;同时,许多计算与结构研究往往未把B2M的动态参与、糖基化差异与氧化还原敏感性纳入统一框架,提示未来需进一步整合结构学与功能免疫学证据以明确其非经典免疫调节潜能 [1,2,3]。
2. 可溶性B2M的非免疫效应与生物标志物价值:优势与局限
在免疫功能之外,血源性B2M在中枢神经系统中的作用提供了较为连续的“关联—干预—机制”证据链:研究通过连体动物、血浆输注与系统给药等范式,将唐氏综合征(DS)血浆中升高的B2M与突触缺陷和记忆损害联系起来,并用B2m基因缺失或抗-B2M抗体中和缓解表型;进一步指出B2M可与NMDAR亚基GluN1的S2环结合并拮抗NMDAR功能,竞争性短肽阻断后可恢复NMDAR依赖的突触功能,从而将B2M扩展为一种内源性NMDAR拮抗剂并提示其在认知障碍中的直接致病潜力 [4]。同时,仍需谨慎外推:系统给药浓度与人类疾病水平的可比性、B2M影响脑内受体的动力学路径,以及结合位点的结构/亲和力证据仍有限,这些因素限制了直接转化推断 [4]。
从生物标志物角度,B2M具有跨疾病预测潜力,但特异性与因果解释存在挑战。一般人群与非CKD人群的大队列数据显示,B2M与全因死亡率及心血管死亡风险呈J形非线性相关,提示其可能整合了慢性炎症、免疫激活与肾清除等多维信息,具有一定风险分层价值 [5]。但观测性研究难以排除残余混杂,且阈值确定与背景校正是临床应用的关键限制 [5]。在CKD场景中,无抗体MRM质谱平台将B2M纳入多标志物面板进行绝对定量,显示与ELISA相关性良好、与eGFR显著相关,并在联合分析中提升对不良结局的预测能力,提示以标准化、多重化量化策略可部分克服单一指标的局限 [6]。与此同时,由于B2M强受肾清除影响,其在非肾病背景下的解读需严格区分“肾功能下降导致的升高”与“炎症/免疫激活信号”的差异 [6]。
3. B2M-MHC-I抗原呈递机制与调控:从装配到免疫逃逸
成熟MHC-I由重链、B2M与结合肽组成,B2M通过非共价相互作用稳定重链折叠并促进复合体转运至细胞表面;B2M缺失/下调会导致表面MHC-I表达丢失并削弱CD8+ T细胞识别,形成免疫逃逸的直接基础 [7,8]。抗原加工与呈递系统(APM)还依赖蛋白酶体亚单位(如PSMB8/9/10)产肽、TAP1/2转运以及ERAP修饰等环节,多个组分协同缺陷会显著降低新生肽-MHC-I复合体形成,并与免疫治疗抵抗相关 [8,9]。功能研究进一步支持因果性:双等位基因丧失或CRISPR敲除B2m可在体内产生对PD-1阻断的耐药表型,提示APM破坏在特定情境下足以驱动ICI失败 [7]。
在调控层面,炎性信号可增强APM转录:Type I干扰素通过STAT1/STAT2-IRF7轴与转录辅激活因子(含NLRC5)上调MHC-I、TAP1、PSMB8及B2M等,促进抗原呈递与T细胞激活 [10,11]。相反,TGF-β可经ALK5-Smad3通路下调多种APM基因,且存在时间学差异(如ERAP1较早下降、B2M更晚下调),提示慢性免疫抑制微环境可能由此逐步形成 [9]。此外,EZH2可在HLA基因以及B2M、TAP1启动子处引入H3K27me3实现持续沉默,并提示EZH2抑制可能恢复APM表达、增强免疫清除潜力 [12]。但需要强调,跨肿瘤类型的异质性与替代免疫路径会修正“B2M缺失=必然无效”的简单推断:例如在MMRd肿瘤中,尽管B2M双等位基因失活常见,ICI仍可能通过CD4+ T细胞效应获得抗肿瘤活性 [13];此外,包含B2M或HLA突变的肿瘤往往伴随更高潜在新抗原负荷,但其是否能通过非经典路径转化为有效免疫识别仍缺乏充分功能证据 [14]。
4. B2M相关信号通路:免疫细胞互作与肿瘤内源性轴
在肿瘤微环境中,B2M状态与多条信号轴共同塑造免疫抑制格局。研究显示,肿瘤相关NK细胞可驱动髓系抑制性细胞(MDSC)表型增强,并与IL-6信号及MDSC标志物(如S100A8/9、ARG1)相关;在动物模型中干预IL-6/STAT3轴可缓解T细胞抑制并抑制肿瘤生长,将NK-MDSC互作的关键节点集中于IL-6/STAT3通路 [15]。与此同时,关于“接触依赖性受体”与“分泌因子”各自贡献仍缺少定量拆分证据,这限制了跨肿瘤类型推广的可预测性 [15]。
B2M缺失对NK细胞内在信号也可能产生直接影响:在NK-92MI细胞系中敲除B2M后,激活受体表达、细胞因子产生与细胞毒性下降,并伴随SHP-1表达及磷酸化上调,从而抑制STAT3与ERK磷酸化;敲除PTPN6(SHP-1)可部分恢复细胞毒性,而CAR改造可进一步克服低反应性,提示SHP-1介导的去磷酸化是抑制节点之一 [16]。但该结论主要基于细胞系,向初代NK许可机制外推仍需谨慎,且SHP-1并非wei一负调控因素 [16]。
除免疫互作外,B2M亦可参与肿瘤细胞内源性信号:在胶质母细胞瘤中,B2M与PIP5K1A互作可激活PI3K/AKT/mTOR并促进MYC介导的TGFβ1分泌,进而诱导巨噬细胞M2样极化,支持肿瘤干性与免疫抑制微环境 [17]。因此,对B2M功能的概括需显式纳入组织与细胞谱系差异 [17]。
5. B2M与疾病:免疫逃逸、微环境、监测与非肿瘤病理
5.1 肿瘤中的B2M缺失:免疫逃逸的基因组学证据
大规模结直肠癌基因组研究显示,MSI-high肿瘤常出现抗原呈递机器的免疫编辑特征,包括B2M与HLA基因的双等位基因丧失(含拷贝数改变与拷贝中性LOH)[18]。临床队列进一步报告MSI-H CRC中B2M突变率可达约24%,且突变与蛋白表达缺失高度一致,支持“基因型—表型”的紧密关联 [19];同时,B2M热点突变在微卫星不稳定CRC中富集,且可能在免疫治疗前已存在,提示其既可能是早期演化事件,也可能在治疗压力下被选择 [20]。
在其他肿瘤中,B2M改变与HLA表达降低、CD8+ T细胞浸润减少等表型共同出现,强化了“抗原呈递受损→免疫逃逸”的解释框架,例如EBV阳性DLBCL中B2M突变与HLA表达降低及CD8+浸润减少并存 [21],并与多类EBV相关或血液肿瘤中HLA相关基因频繁受损的模式一致 [22]。在恶性胶质瘤中,针对HLA区域测序复杂性的研究亦识别出B2M与TAP基因体细胞突变,并提示其在复发GBM中可能富集,指向进展或治疗过程中选择压力的作用 [23]。在MSI/dMMR胰腺导管腺癌中,B2M失活可表现出原发与转移灶间异质性,提示治疗评估需关注空间差异与转移部位的重要性 [24]。
5.2 B2M、肿瘤微环境与预后:情景依赖的免疫表型
在DLBCL中,B2M等基因突变与多处结外侵及(ENI)、更差预后以及TME层面的Treg招募增强、基质基因表达降低相关;动物模型亦提示B2M突变与骨髓侵及相关,暗示其可能通过抗原呈递下降与免疫浸润谱改变共同驱动侵袭表型 [25]。但目前对其如何直接驱动Treg募集(如趋化网络或基质改造)仍以相关性为主。
在林奇综合征相关CRC中,即便存在肿瘤细胞B2M丧失,仍可观察到PD-L1上调与丰富的CD3+/CD8+浸润,提示B2M缺失并不必然等同于“完全免疫隐匿”,应结合更广泛免疫景观解释ICI敏感性 [26]。此外,PMBL整合基因组研究显示包括B2M在内的多种免疫逃逸相关基因频繁突变并与JAK-STAT、NF-κB改变共存,支持免疫回避为该亚型核心机制之一 [27];滤泡淋巴瘤单细胞研究亦在Treg细胞中识别出与免疫检查点共表达网络相关的基因集,包含B2M,提示其在免疫细胞内源性表达与调控中可能具有意义,但仍缺乏直接功能验证 [28]。
5.3 B2M/HLA基因组学与动态监测:从起源到MRD
cHL对HRS细胞分选后进行全基因组测序揭示复杂结构变异与演化时序:包括B2M等免疫逃逸相关基因的突变往往发生在大范围染色体增益之前,从时间尺度上支持其为早期关键事件 [29]。在更可及的临床监测层面,针对ctDNA的快速基因面板研究显示诊断时总体变异检出率较高,并报告B2M突变率约33.3%;治疗早期ctDNA迅速转阴并与PET指标高度一致,提示其可作为动态生物标志物与影像互补 [30]。但“不可检测”受到检测限、肿瘤DNA释放量与面板覆盖范围限制,MRD与克隆演化监测仍可能存在盲区。
此外,B2M也出现在DLBCL遗传分型中(如EZB-like亚型常见变异之一),提示其跨淋巴瘤亚型具有一定普遍性,并为分型与免疫相关决策提供潜在线索 [31]。
5.4 非肿瘤疾病:组织与样本类型依赖的B2M异常
在pSS唾液腺微阵列差异表达与网络分析中,B2M显著上调且位列枢纽基因,上调基因富集于抗原呈递与干扰素通路,提示其与局部免疫激活密切相关,但仍需功能验证区分驱动与伴随标志 [32]。RA外周血单细胞研究则在DNT细胞群体中观察到包括B2M在内的多基因下调,并伴随氧化磷酸化受抑与抗原呈递缺陷,提示在某些免疫细胞亚群中B2M降低可能与代谢与呈递功能下降并行,但转录与蛋白/表面表达的一致性仍需实验确认 [33]。KD牙龈沟液蛋白组学研究鉴定B2M等上调蛋白并提示其作为早期诊断标志物的可能性,但局部体液来源与系统性病理之间的因果路径仍待厘清 [34]。此外,透析可显著降低循环B2M水平,提示在肾功能受损或接受透析干预的人群中,B2M变化可能强受治疗操作影响,构成重要混淆因素 [35]。
6. B2M靶向药物最近研究进展
靶向β2微球蛋白(B2M)的药物研发近年来呈现多元化趋势,适应症从传统的肿瘤免疫治疗拓展至自身免疫性疾病及神经退行性疾病。目前进展最快的项目聚焦于黑色素瘤及糖尿病等领域,同时基于基因编辑和通用型CAR-T技术的创新疗法也进入临床验证阶段,显示出B2M作为多功能靶点的转化潜力。部分在研管线整理如下表:
| 药物 | 类型 | 适应症 | 研发阶段 | 研发机构 |
|---|---|---|---|---|
| B2M单抗 | 单克隆抗体 | 黑色素瘤 | 临床前 | 多个研究机构 |
| B2M抑制剂 | 小分子化合物 | 1型糖尿病 | 临床前 | 多家机构 |
| B2M基因编辑CAR-T | 细胞治疗 | 血液肿瘤 | 临床1期 | 多个临床中心 |
| 通用型CAR-T(B2M敲除) | 细胞治疗 | 实体瘤/血液肿瘤 | 临床1/2期 | 生物技术公司 |
| B2M中和抗体 | 抗体药物 | 阿尔茨海默病 | 探索阶段 | 科研机构 |
| B2M疫苗 | 疫苗 | 肿瘤免疫治疗 | 临床前 | 药企合作项目 |
7. B2M研究工具推荐:重组蛋白、抗体与ELISA试剂盒选型指南
B2M在“抗原呈递—免疫逃逸—微环境重塑—临床监测—治疗工程化”链条中占据关键节点:其缺失或突变可通过MHC-I装配受损削弱CD8+ T细胞识别,并在多肿瘤类型中与免疫治疗耐药相关。华美生物提供B2M重组蛋白、抗体及ELISA试剂盒产品,助力您进行相关机制研究及靶向药物开发。
● B2M 重组蛋白
Recombinant Human HLA-A*02:01&B2M&NY-ESO-1 (SLLMWITQV) protein, Biotinylated
CSB-CSB-MP6862HU(M1)-B
Recombinant Sheep Beta-2-microglobulin (B2M)
CSB-EP002486SH
● B2M 抗体
B2M Recombinant Monoclonal Antibody
CSB-RA945176A0HU
B2M Monoclonal Antibody
CSB-MA057861A0m
● B2M ELISA 试剂盒
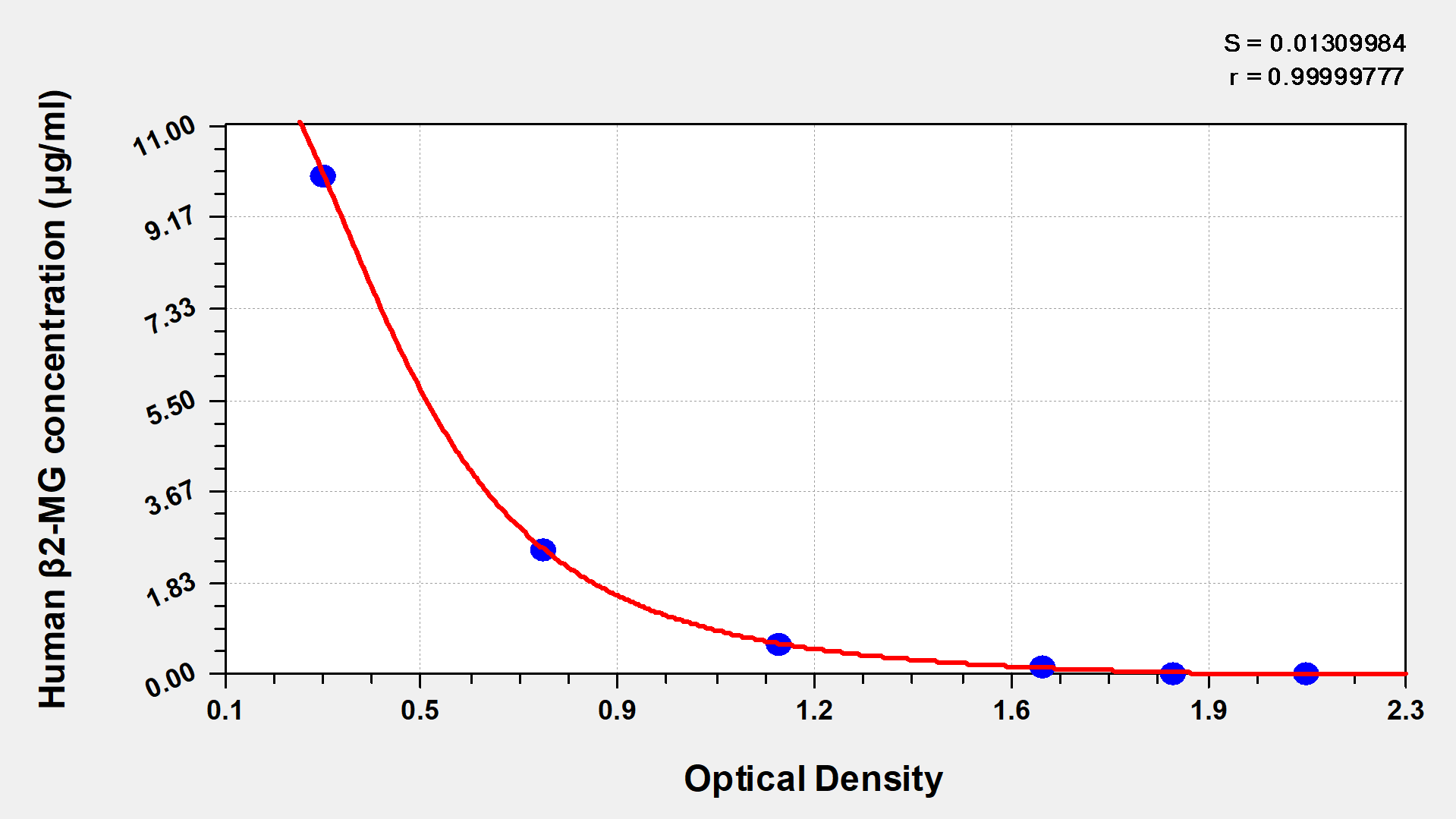
Human β2-microglobulin,BMG/β2-MG ELISA Kit
CSB-E04883h
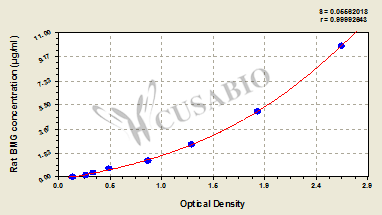
Rat β2-microglobulin,BMG/β2-MG ELISA Kit
CSB-E11298r
[1] Hans-Christof Gasser, Georges Bedran, Bo Ren, David Goodlett, Javier Alfaro, Ajitha Rajan.(2021). Interpreting BERT architecture predictions for peptide presentation by MHC class I proteins.
[2] J. C. Phillips.(2022). Evolution and Mutations of Beta 2 Microglobulin.
[3] Mepur H Ravindranath, Narendranath M Ravindranath, Senthamil R Selvan, Fatiha El Hilali, Carly J Amato-Menker, Edward J Filippone.(2023). Cell Surface B2m-Free Human Leukocyte Antigen (HLA) Monomers and Dimers: Are They Neo-HLA Class and Proto-HLA?.
[4] Yue Gao, Yujuan Hong, Lihong Huang, Shuang Zheng, Haibin Zhang, Shihua Wang, Yi Yao, Yini Zhao, Lin Zhu, Qiang Xu, Xuhui Chai, Yuanyuan Zeng, Yuzhe Zeng, Liangkai Zheng, Yulin Zhou, Hong Luo, Xian Zhang, Hongfeng Zhang, Ying Zhou, Guo Fu, Hao Sun, Timothy Y Huang, Qiuyang Zheng, Huaxi Xu, Xin Wang.(2023). β2-microglobulin functions as an endogenous NMDAR antagonist to impair synaptic function..
[5] Hang Fang, Qiankun Zhang, Lie Jin.(2023). Association of beta-2-microglobulin with cardiovascular and all-cause mortality in the general and non-CKD population..
[6] Manousos Makridakis, Georgia Kontostathi, Eleni Petra, Rafael Stroggilos, Vasiliki Lygirou, Szymon Filip, Flore Duranton, Harald Mischak, Angel Argiles, Jerome Zoidakis, Antonia Vlahou.(2020). Multiplexed MRM-based protein quantification of putative prognostic biomarkers for chronic kidney disease progression in plasma..
[7] Scott Gettinger, Jungmin Choi, Katherine Hastings, Anna Truini, Ila Datar, Ryan Sowell, Anna Wurtz, Weilai Dong, Guoping Cai, Mary Ann Melnick, Victor Y. Du, Joseph Schlessinger, Sarah B. Goldberg, Anne C. Chiang, Miguel F. Sanmamed, Ignacio Melero, Jackeline Agorreta, Luis M. Montuenga, Richard P. Lifton, Soldano Ferrone, Paula Kavathas, David L. Rimm, Susan M. Kaech, Kurt A. Schalper, Roy S. Herbst, Katerina Politi.(2017). Impaired HLA Class I Antigen Processing and Presentation as a Mechanism of Acquired Resistance to Immune Checkpoint Inhibitors in Lung Cancer.
[8] Karolina Žilionytė, Ugnė Bagdzevičiūtė, Agata Mlynska, Elena Urbštaitė, Emilija Paberalė, Neringa Dobrovolskienė, Jan Aleksander Krasko, Vita Pašukonienė.(2022). Functional antigen processing and presentation mechanism as a prerequisite factor of response to treatment with dendritic cell vaccines and anti-PD-1 in preclinical murine LLC1 and GL261 tumor models..
[9] Alix K. Berglund, Anna L. Hinson, Lauren V. Schnabel.(2023). TGF-β downregulates antigen processing and presentation genes and MHC I surface expression through a Smad3-dependent mechanism.
[10] Hemin Jiang, Yue Li, Min Shen, Yucheng Liang, Qian Yu, Hao Dai, Kuanfeng Xu, Xinyu Xu, Hui Lv, Jie Zhang, Tao Yang, Qi Fu.(2022). Interferon‐α promotes MHC I antigen presentation of islet β cells through STAT1‐IRF7 pathway in type 1 diabetes.
[11] Chrissie E B Ong, Amanda L Patchett, Jocelyn M Darby, Jinying Chen, Guei-Sheung Liu, A Bruce Lyons, Gregory M Woods, Andrew S Flies.(2021). NLRC5 regulates expression of MHC-I and provides a target for anti-tumor immunity in transmissible cancers..
[12] Wei Ding, Zihan Ding, Qinghong Zeng, Yan Qiu, Christopher R Donnelly, Yuqi Wu, Yuchen Jiang, Qi Han, Hao Xu, Hao Cui, Xiangfei Liu, Xin Chen, Sixin Jiang, Mei Huang, Dan Pan, Dan Yang, Li Li, Lihong Yao, Minghai Tang, Jing Li, Taiwen Li, Xiaoping Xu, Qianming Chen, Hang Zhao, Longyu Li, Lu Jiang, Xiaobo Luo.(2025). Prevention of cancer initiation by augmenting MHC-I antigen presentation via EZH2 inhibition..
[13] Giovanni Germano, Steve Lu, Giuseppe Rospo, Simona Lamba, Benoı̂t Rousseau, Sonia Fanelli, Denise Stenech, Dung T. Le, John L. Hays, Maria Grazia Totaro, Vito Amodio, Rosaria Chilà, Anna Mondino, Luis A. Díaz, Federica Di Nicolantonio, Alberto Bardelli.(2021). CD4 T Cell–Dependent Rejection of Beta-2 Microglobulin Null Mismatch Repair–Deficient Tumors.
[14] Andrea Castro, Kıvılcım Öztürk, Rachel Marty Pyke, Xian Su, Maurizio Zanetti, Hannah Carter.(2019). Elevated neoantigen levels in tumors with somatic mutations in the HLA-A, HLA-B, HLA-C and B2M genes.
[15] Shiyong Neo, Le Tong, Joni Chong, Yaxuan Liu, Jing Xu, Mariana M.S. Oliveira, Yi Chen, Ziqing Chen, Keene Lee, Nutsa Burduli, Xinsong Chen, Juan Gao, Ran Ma, Jia Pei Lim, Jianxin Huo, Shengli Xu, Evren Alici, Stina L. Wickström, Felix Haglund, Johan Hartman, Arnika Kathleen Wagner, Yihai Cao, Rolf Kiessling, Kong‐Peng Lam, Lisa S. Westerberg, Andreas Lundqvist.(2024). Tumor-associated NK cells drive MDSC-mediated tumor immune tolerance through the IL-6/STAT3 axis.
[16] Kuo Yu, Xiaolong Liu, Guangyuan Wu, Zhongyao An, Xin Wang, Yang Liu, Hailong Wang, Mingli Huang, Linlin Zhao, Ce Shi, Xin Sun, Lu Xu, Sen Qi, Xin Zhang, Yueqiu Teng, Song Guo Zheng, Zhiren Zhang, Zhenkun Wang.(2025). Targeting SHP-1-Mediated Inhibition of STAT3 and ERK Signalling Pathways Rescues the Hyporesponsiveness of MHC-I-Deficient NK-92MI..
[17] Daqi Li, Qian Zhang, Lu Li, Kexin Chen, Junlei Yang, Deobrat Dixit, Ryan C Gimple, Shusheng Ci, Chenfei Lu, Lang Hu, Jiancheng Gao, Danyang Shan, Yangqing Li, Junxia Zhang, Zhumei Shi, Danling Gu, Wei Yuan, Qiulian Wu, Kailin Yang, Linjie Zhao, Zhixin Qiu, Deguan Lv, Wei Gao, Hui Yang, Fan Lin, Qianghu Wang, Jianghong Man, Chaojun Li, Weiwei Tao, Sameer Agnihotri, Xu Qian, Yu Shi, Yongping You, Nu Zhang, Jeremy N Rich, Xiuxing Wang.(2022). β2-Microglobulin Maintains Glioblastoma Stem Cells and Induces M2-like Polarization of Tumor-Associated Macrophages..
[18] Catherine S. Grasso, Marios Giannakis, Daniel K. Wells, Tsuyoshi Hamada, Xinmeng Jasmine Mu, Michael J. Quist, Jonathan A. Nowak, Reiko Nishihara, Zhi Rong Qian, Kentaro Inamura, Teppei Morikawa, Katsuhiko Nosho, Gabriel Abril-Rodríguez, Charles M. Connolly, Helena Escuin-Ordinas, Milan S. Geybels, William M. Grady, Li Hsu, Siwen Hu‐Lieskovan, Jeroen R. Huyghe, Yeon Joo Kim, Paige Krystofinski, Mark D.M. Leiserson, Dennis Montoya, Brian B. Nadel, Matteo Pellegrini, Colin C. Pritchard, Cristina Puig-Saus, Elleanor H. Quist, Benjamin J. Raphael, Stephen J. Salipante, Daniel Sanghoon Shin, Eve Shinbrot, Brian H. Shirts, Sachet A. Shukla, Janet L. Stanford, Wei Sun, Jennifer Tsoi, Alexander Upfill‐Brown, David A. Wheeler, Catherine J. Wu, Ming Yu, Syed Hassan Ejaz Zaidi, Jesse M. Zaretsky, Stacey Gabriel, Eric S. Lander, Levi A. Garraway, Thomas J. Hudson, Charles S. Fuchs, Antoni Ribas, Shuji Ogino, Ulrike Peters.(2018). Genetic Mechanisms of Immune Evasion in Colorectal Cancer.
[19] Sumit Middha, Rona Yaeger, Jinru Shia, Zsofia K. Stadler, Sarah King, Shanna Guercio, Viktoriya Paroder, David D. B. Bates, Satshil Rana, Luis A. Díaz, Leonard B. Saltz, Neil H. Segal, Marc Ladanyi, Ahmet Zehir, Jaclyn F. Hechtman.(2019). Majority ofB2M-Mutant and -Deficient Colorectal Carcinomas Achieve Clinical Benefit From Immune Checkpoint Inhibitor Therapy and Are Microsatellite Instability-High.
[20] Su Yeon Yeon, Seung-Hyun Jung, Yun Sol Jo, Eun Ji Choi, Min Sung Kim, Yeun-Jun Chung, Sug Hyung Lee.(2019). Immune checkpoint blockade resistance-related B2M hotspot mutations in microsatellite-unstable colorectal carcinoma..
[21] Miaoxia He, Bin Liu, Gusheng Tang, Lijuan Jiao, Xuefei Liu, Shuyi Yin, Tao Wang, Jie Chen, Lei Gao, Xiong Ni, Libin Wang, Lili Xu, Jianmin Yang.(2022). B2M mutation paves the way for immune tolerance in pathogenesis of Epstein-Barr virus positive diffuse large B-cell lymphomas..
[22] Chantana Polprasert, Yasuhide Takeuchi, Hideki Makishima, Kitsada Wudhikarn, Nobuyuki Kakiuchi, Nichthida Tangnuntachai, Thamathorn Assanasen, Wimonmas Sitthi, Hamidah Muhamad, Panisinee Lawasut, Sunisa Kongkiatkamon, Udomsak Bunworasate, Koji Izutsu, Yuichi Shiraishi, Kenichi Chiba, Hiroko Tanaka, Satoru Miyano, Seishi Ogawa, Kenichi Yoshida, Ponlapat Rojnuckarin.(2020). Frequent mutations in HLA and related genes in extranodal NK/T cell lymphomas.
[23] Sara C Schulte, Wolfgang Peter, Georg Rosenberger, Moritz Schäfer, Cecile L Maire, Alessandra Rünger, Alice Ryba, Kristoffer Riecken, Krystian D Fita, Jakob Matschke, Nuray Akyüz, Judith Dierlamm, Gunnar W Klau, Franz L Ricklefs, Jens Gempt, Manfred Westphal, Katrin Lamszus, Alexander Dilthey, Malte Mohme.(2025). Somatic Mutations in HLA Class Genes and Antigen-Presenting Molecules in Malignant Glioma..
[24] Claudio Luchini, Andrea Mafficini, Deyali Chatterjee, Maria L Piredda, Concetta Sciammarella, Pooja Navale, Giuseppe Malleo, Paola Mattiolo, Giovanni Marchegiani, Antonio Pea, Roberto Salvia, Lodewijk A Brosens, Gaetano Paolino, Maria G Mastrosimini, Nicola Silvestris, Michele Milella, Liang Cheng, Volkan N Adsay, Rita T Lawlor, Aldo Scarpa.(2022). Histo-molecular characterization of pancreatic cancer with microsatellite instability: intra-tumor heterogeneity, B2M inactivation, and the importance of metastatic sites..
[25] Rong Shen, Pengpeng Xu, Nan Wang, Hongmei Yi, Lei Dong, Di Fu, Jinyan Huang, Hengye Huang, Anne Janin, Shu Cheng, Li Wang, Weili Zhao.(2020). Influence of oncogenic mutations and tumor microenvironment alterations on extranodal invasion in diffuse large B‐cell lymphoma.
[26] Joanna Walkowska, Thomas Kallemose, Göran Jönsson, Mats Jönsson, Ove Andersen, Mads Hald Andersen, Inge Marie Svane, Anne Langkilde, Mef Nilbert, Christina Therkildsen.(2018). Immunoprofiles of colorectal cancer from Lynch syndrome.
[27] Anja Mottok, Stacy Hung, Elizabeth A. Chavez, Bruce W. Woolcock, Adèle Telenius, Lauren C. Chong, Barbara Meissner, Hisae Nakamura, Christopher Rushton, Elena Viganò, Clémentine Sarkozy, Randy D. Gascoyne, Joseph M. Connors, Susana Ben‐Neriah, Andrew J. Mungall, Marco A. Marra, Reiner Siebert, David W. Scott, Kerry J. Savage, Christian Steidl.(2019). Integrative genomic analysis identifies key pathogenic mechanisms in primary mediastinal large B-cell lymphoma.
[28] Noemi Andor, Erin F. Simonds, Debra K. Czerwinski, Jiamin Chen, Susan M. Grimes, Christina Wood-Bouwens, Grace Zheng, Matthew Kubit, Stephanie Greer, William A. Weiss, Ronald Levy, Hanlee P. Ji.(2018). Single-cell RNA-Seq of follicular lymphoma reveals malignant B-cell types and coexpression of T-cell immune checkpoints.
[29] Francesco Maura, Bachisio Ziccheddu, Jenny Xiang, Bhavneet Bhinder, Joel Rosiene, Federico Abascal, Kylee Maclachlan, Kenneth Wha Eng, Manik Uppal, Feng He, Wei Zhang, Qi Gao, Venkata D. Yellapantula, Vicenta Trujillo-Alonso, Sunita I. Park, Matthew J. Oberley, Elizabeth Ruckdeschel, Megan S. Lim, Gerald Wertheim, Matthew J. Barth, Terzah M. Horton, Andriy Derkach, Alexandra E. Kovach, Christopher J. Forlenza, Yanming Zhang, Ola Landgren, Craig H. Moskowitz, Ethel Cesarman, Marcin Imieliński, Olivier Elemento, Mikhail Roshal, Lisa Giulino‐Roth.(2023). Molecular Evolution of Classic Hodgkin Lymphoma Revealed Through Whole-Genome Sequencing of Hodgkin and Reed Sternberg Cells.
[30] Vincent Camus, Mathieu Viennot, Justine Lequesne, Pierre‐Julien Viailly, Élodie Bohers, Lucile Bessi, B. Marcq, Pascaline Etancelin, Sydney Dubois, Jean‐Michel Picquenot, Elena‐Liana Veresezan, Marie Cornic, Lucie Burel, Justine Loret, Stéphanie Becker, Pierre Decazes, Pascal Lenain, Stéphane Leprêtre, Emilie Lemasle, Hélène Lanic, Anne‐Lise Ménard, Nathalie Contentin, Hervé Tilly, Aspasia Stamatoullas, Fabrice Jardin.(2020). Targeted genotyping of circulating tumor DNA for classical Hodgkin lymphoma monitoring: a prospective study.
[31] Rong Shen, Di Fu, Lei Dong, Mu‐Chen Zhang, Qing Shi, Zi‐Yang Shi, Shu Cheng, Li Wang, Pengpeng Xu, Weili Zhao.(2023). Simplified algorithm for genetic subtyping in diffuse large B-cell lymphoma.
[32] Lei Zhang, Poshi Xu, Xiaoyu Wang, Zongshan Zhang, Wenxin Zhao, Zhengmin Li, Guangxia Yang, Panpan Liu.(2019). Identification of differentially expressed genes in primary Sjögren's syndrome.
[33] Xinqiang Song, Yu Zhang, Lijun Zhao, Jinke Fan, Tao Peng, Ying Ma, Nailiang Guo, Xiaotong Wang, Xudong Liu, Zhe Liu, Lei Wang.(2023). Analyzation of the Peripheral Blood Mononuclear Cells Atlas and Cell Communication of Rheumatoid Arthritis Patients Based on Single-Cell RNA-Seq.
[34] Xue Fan, Ying Li, Yuehao Xu, Jianqing Lin, Xin Guo, Jinwen Liao, Mingguo Xu.(2025). New biomarkers of Kawasaki disease identified by gingival crevicular fluid proteomics..
[35] Jan Kampmann, Michael Mejer Hunderup, Eva Rabing Brix Petersen, Vivi Andersen, Thor Aage Skovsted.(2024). High-sensitive troponin T, suPAR and Beta-2-microglobulin changes in concentration during hemodialysis.



